
Overview
Evolutionary forces have generated enormous amounts of molecular and clinically relevant phenotypic diversity across species, with several independent mammalian lineages evolving long life- and health-spans. An evolutionary lens is a uniquely powerful, orthogonal approach to elucidate natural longevity mechanisms and identify solutions for extended longevity with strong translational potential. However, linking evolutionary sequence changes to target genes, regulatory networks, and biological processes is challenging. Powerful new phylogenetically informed computational approaches able to handle more than 500 mammalian genomes, combined with high-throughput functional genomic screens, present a unique opportunity to decipher the genetic basis of longevity across species.
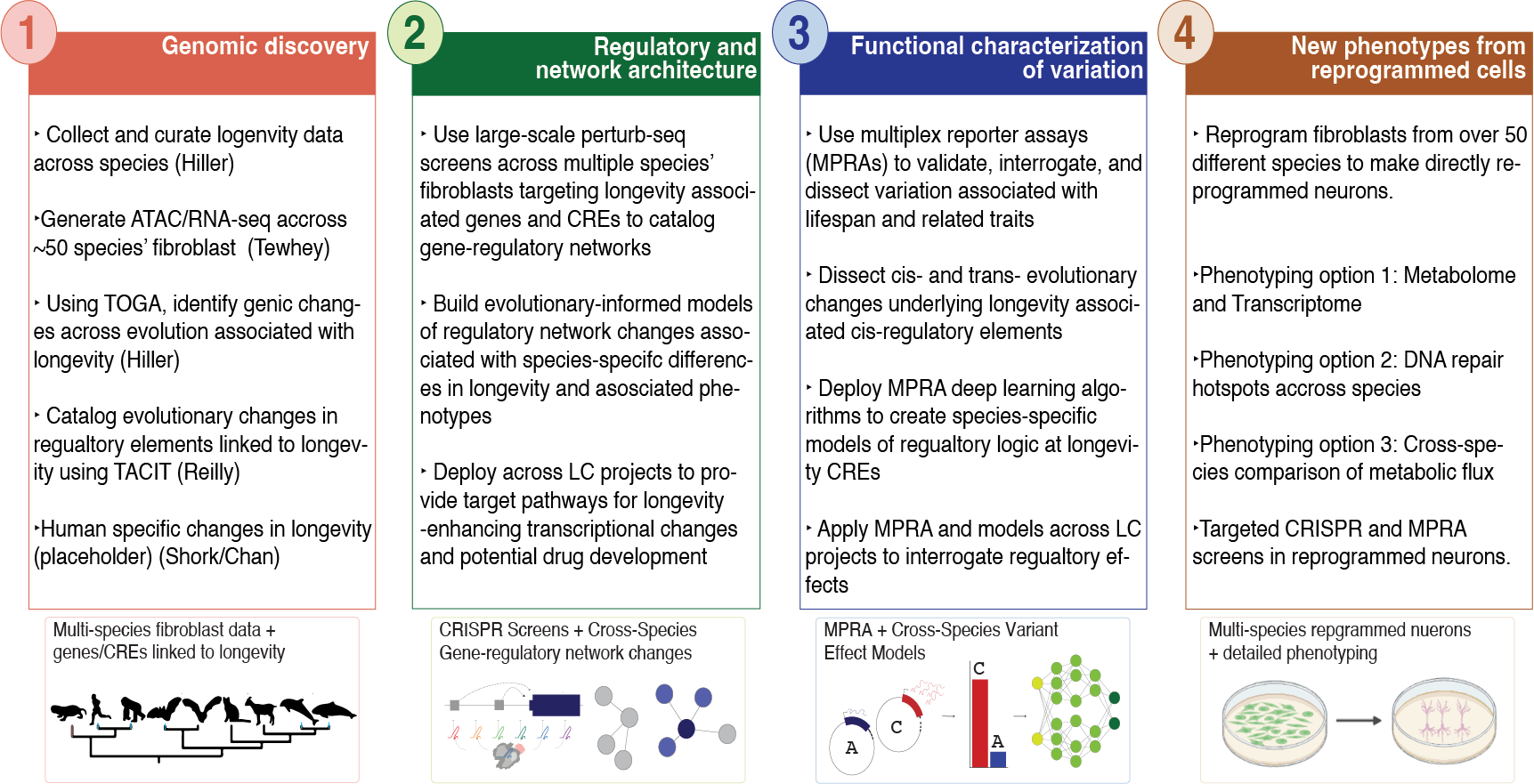
The LC Cross Species and Evolutionary Biology (CSEB) Project will pursue an unprecedented integration across different methodologies and extend to human studies within the LC to evaluate cross-species differences in specific biologic and genetic factors influencing species’ lifespan. By leveraging a large collection of species multiomic data, legacy LC collections of fibroblasts from 158 species, and high-throughput functional assays, we will (i) uncover the genomic basis of extended longevity in mammals to deepen our understanding of aging and reveal new candidates for functional experiments, ii) validate the effects of genes, regulatory elements, and variants associated with longevity across species in humans as targets for geroprotective interventions, iii) address the contribution of abrupt and long-term evolutionary changes leading to human divergence that might explain the uniquely human lifespan and iv) expand relevant induced pluripotent stem cells (iPSCs), neuronal, and other biomaterial collections along with cross-species genomic data models to interrogate longevity in humans.

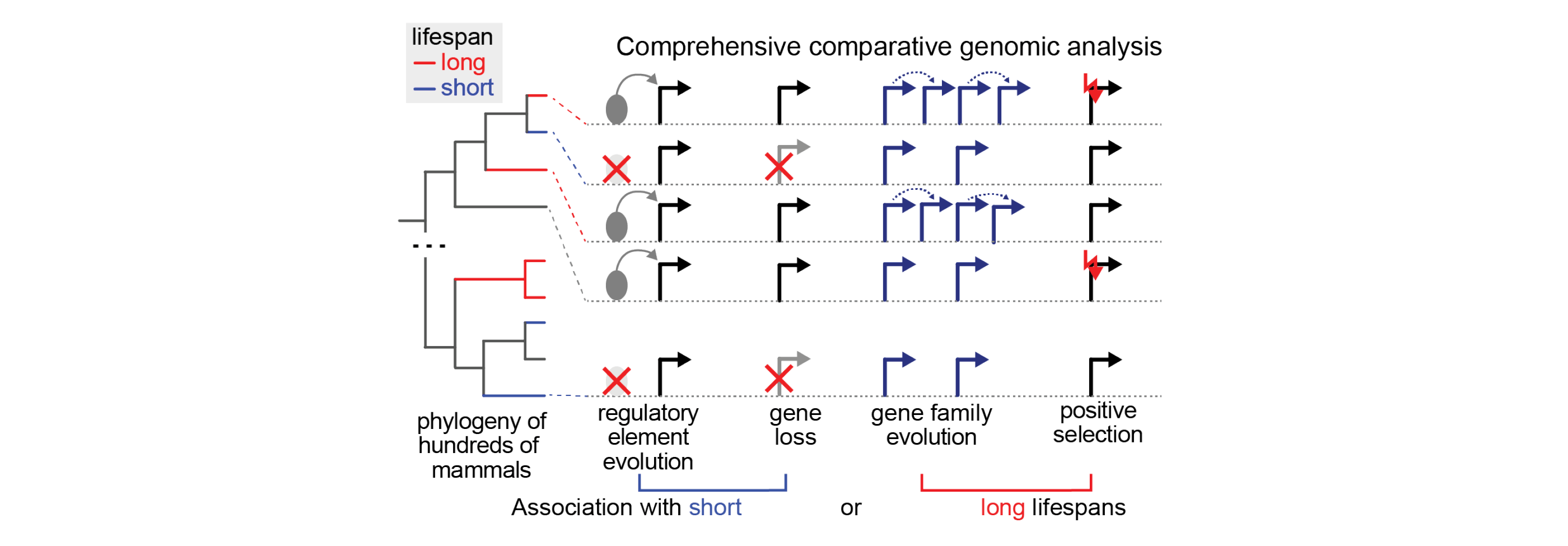
Michael Hiller will co-lead Aim1 “Genomic Discovery”, together with Dr. Chan, to uncover the genomic basis of lifespan differences in mammals. We will mine and curate longevity data and relevant traits from different sources and perform comprehensive comparative genomic analyses, leveraging computational approaches can handle hundreds of genomes. We will systematically screen for selection patterns, gene losses and duplications, and differences in conserved noncoding elements, and associate them with lifespan and related longevity traits using phylogenetic association methods.
Co-Leads
Other Key Personnel
Trainees
